Alphafold Structure Predictions Available In Interpro
interproAlphaFold 2.0 has revolutionised structure prediction enabling the rapid creation of high quality models across many model organisms. We expect these models to drive forward the field of molecular biology and biomedical research.
DeepMind and EMBL’s European Bioinformatics Institute (EMBL-EBI) have launched the AlphaFold Database (AlphaFold DB), a joint project to openly and freely share millions of AlphaFold protein structure predictions with the scientific community. The first AlphaFold DB release contains approximately 375,000 protein structures, covering the proteomes of 22 species, including human, mouse, E. coli and more.
We are delighted to announce that InterPro has connected to the AlphaFold Database of AlphaFold models from its entries so you can now see the structures for all your favourite protein families.
We provide two entry points within InterPro to the AlphaFold structural models. Firstly if you go to the entry for a specific protein and a model is available for that protein you can click on the new AlphaFold tab to view the structure. In the example below we show the rather spectacular structure model of the Arabidopsis SABRE protein.
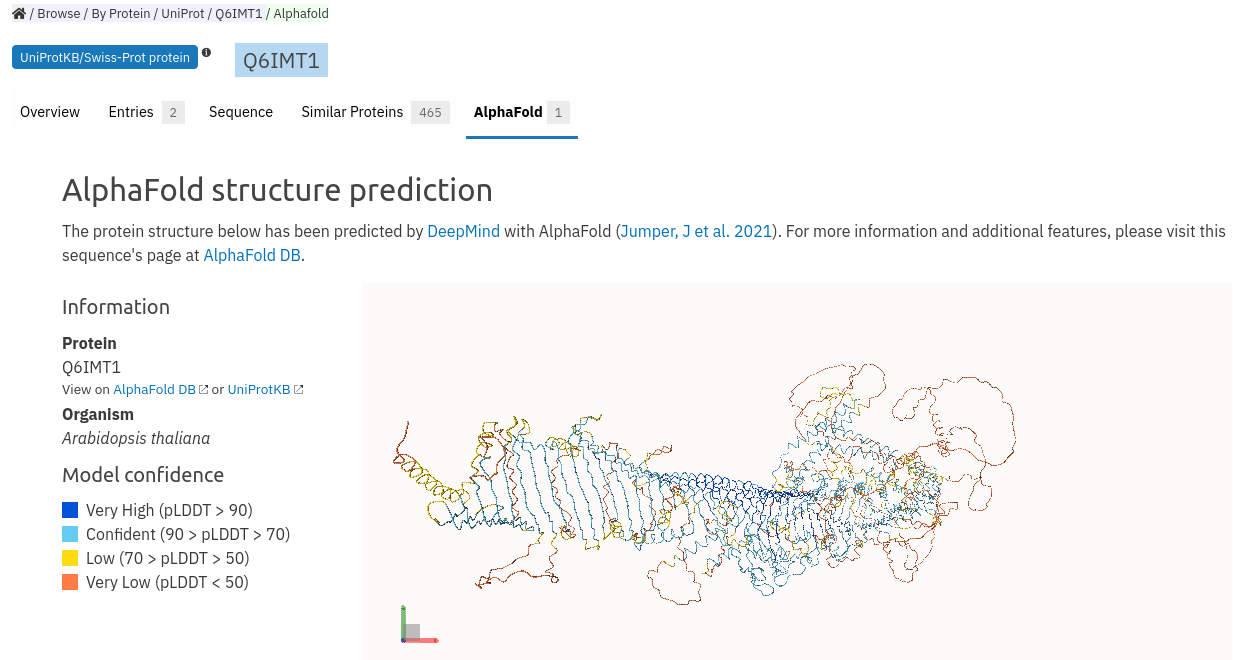
The second entry point is via the InterPro entry pages where you can also access a new AlphaFold tab. By clicking on the tab you will be taken to a page that will show an example AlphaFold model and a table below which shows other models that are available for that entry. In this case there are 19 models available for InterPro entry IPR019441.
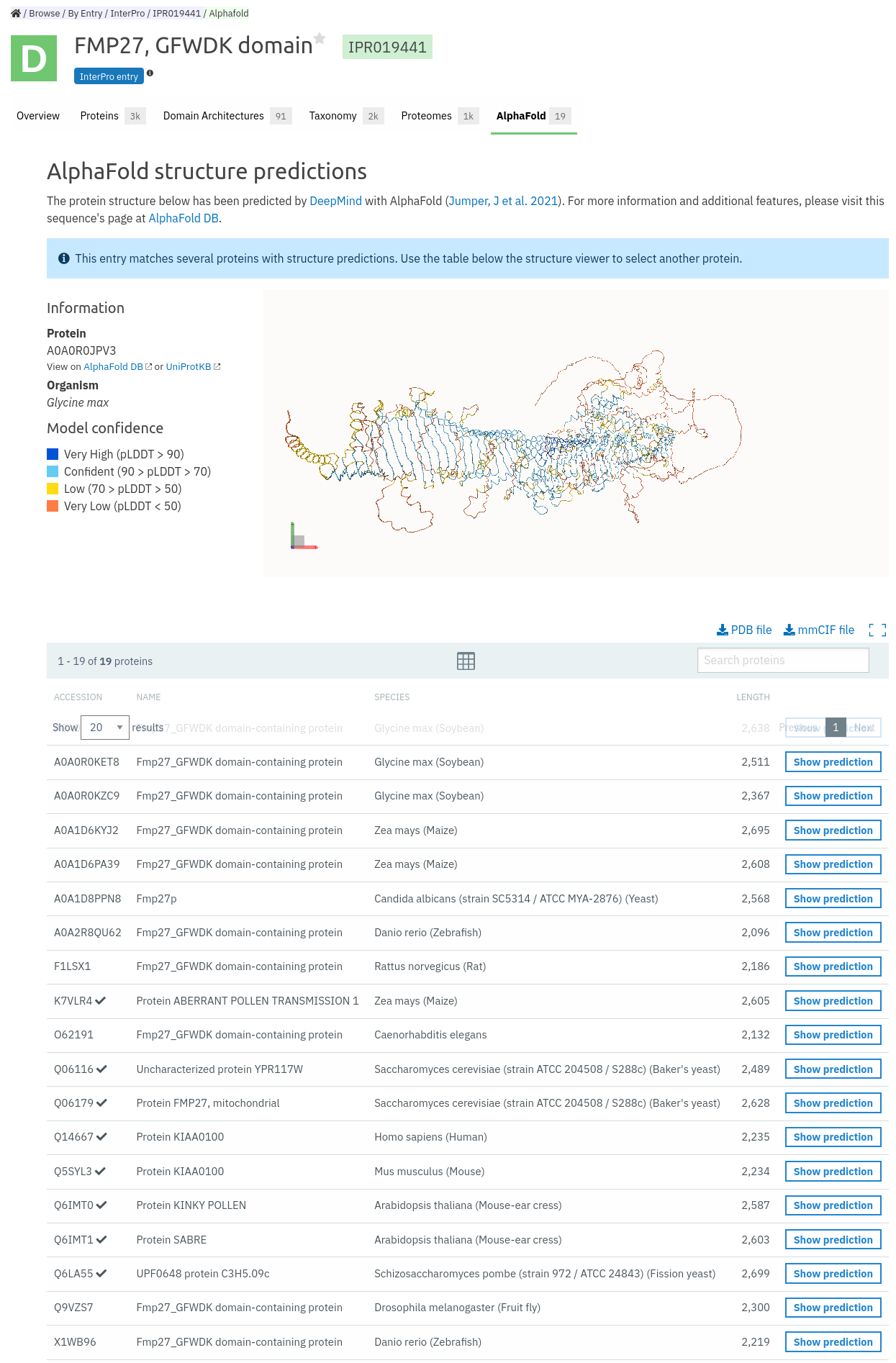
We hope that you are able to make many new exciting discoveries using these new structures and the InterPro website.
